Preparation Technology
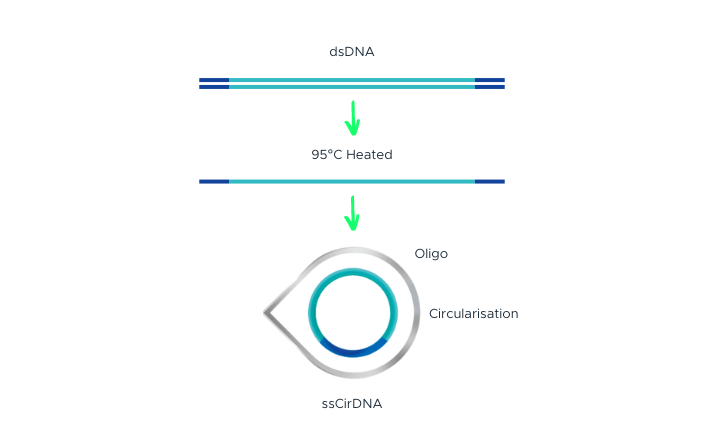
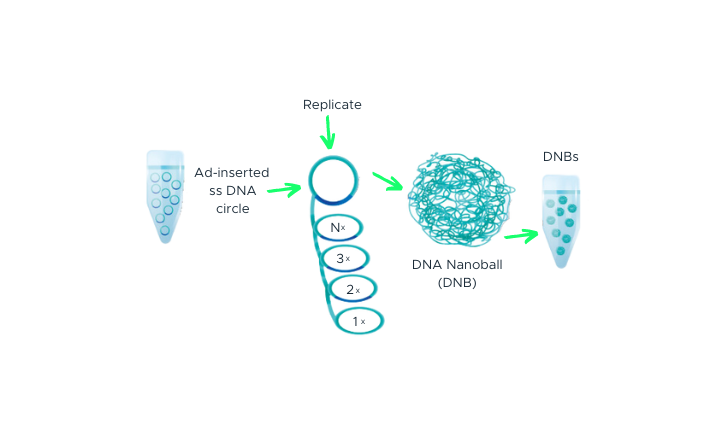
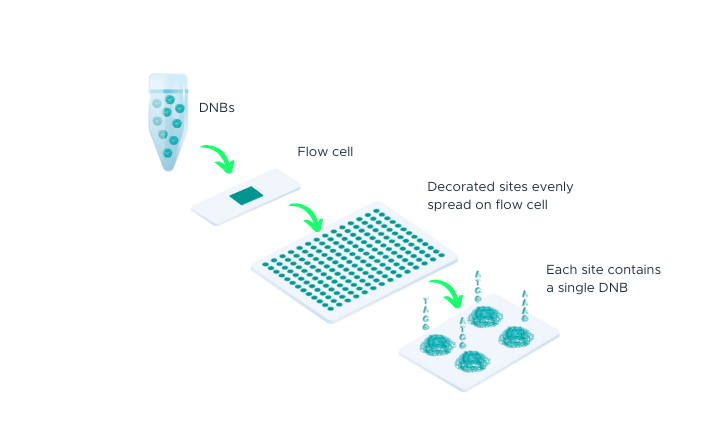
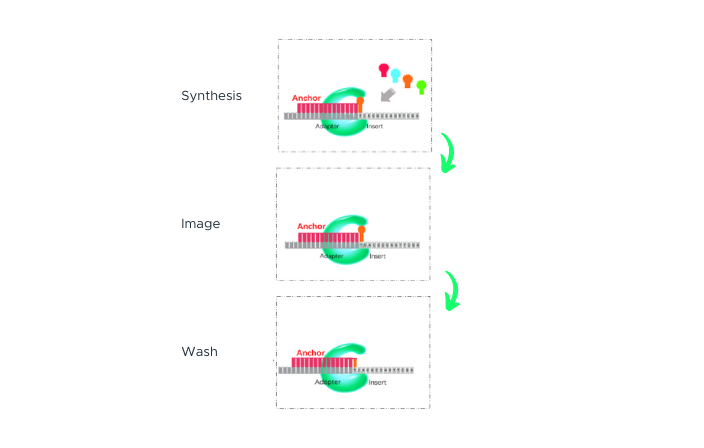
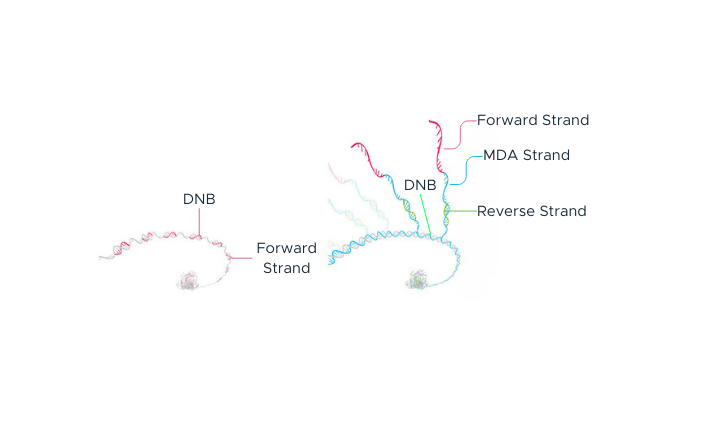
The process begins with double-stranded DNA that contains adapter sequences at both ends. We apply heat to this DNA, causing it to ‘denature’ or separate into two single-stranded DNAs (ssDNA). A specially designed molecule, called a ‘splint oligonucleotide’, plays a crucial role next. This molecule has a sequence that matches both ends of one of the separated strands, allowing it to bind or ‘hybridize’ to these ends. When it does so, it creates a structure known as a ‘nicked circle’. Finally, an enzyme called DNA ligase is used to repair the ‘nick’ in this circle, transforming it into a seamless, single-stranded circle. This carefully controlled process, part of our DNA Nanoball Preparation technology, is one of the reasons you can place your trust in our DNA sequencing capabilities.