

Turbocharge your sequencing - high-speed, high flexibility and ultra-high throughput
Description
Powered by MGI's unique DNBSEQTM Technology, after fully upgrading biochemical, fluidics, and optical systems, DNBSEQ-T7 can output up to 7 Tb of high-quality data within 24 hours and is widely applicable to:
- Whole Genome Sequencing
- Deep Exome Sequencing
- Epigenome Sequencing
- Transcriptome Sequencing
- Tumour Panel
- and other large sequencing projects
Infused with MGI's pioneering DNBSEQTM Technology and bolstered by profound upgrades in biochemical, fluidics, and optical arenas, DNBSEQ-T7 is the ultimate and ultra-high-throughput sequencer for large genome sequencing projects and high-complexity population studies, that can output 1~7TB of high-quality data per day.
- High-velocity Sequencing
Achieve PE150 sequencing in roughly 22 to 24 hours - Adaptable Sequencing Modes
Equipped to handle 4 Flow Cells simultaneously, the DNBSEQ-T7 offers a mix of PE150 and PE100 sequencing options, catering to diverse research demands - Consistent Ultra-high Throughput
Ensuring round-the-clock acquisition of high-fidelity data, expect up to a staggering 7 Tb output daily
*StandardMPS sequencing reagents in modified form are available in Germany, UK, Sweden, and Switzerland.
DNBSEQTM TECHNOLOGY
DNA Nanoball sequencing technology – No accumulation of amplification errors
INTRODUCTION DNBSEQ-T7
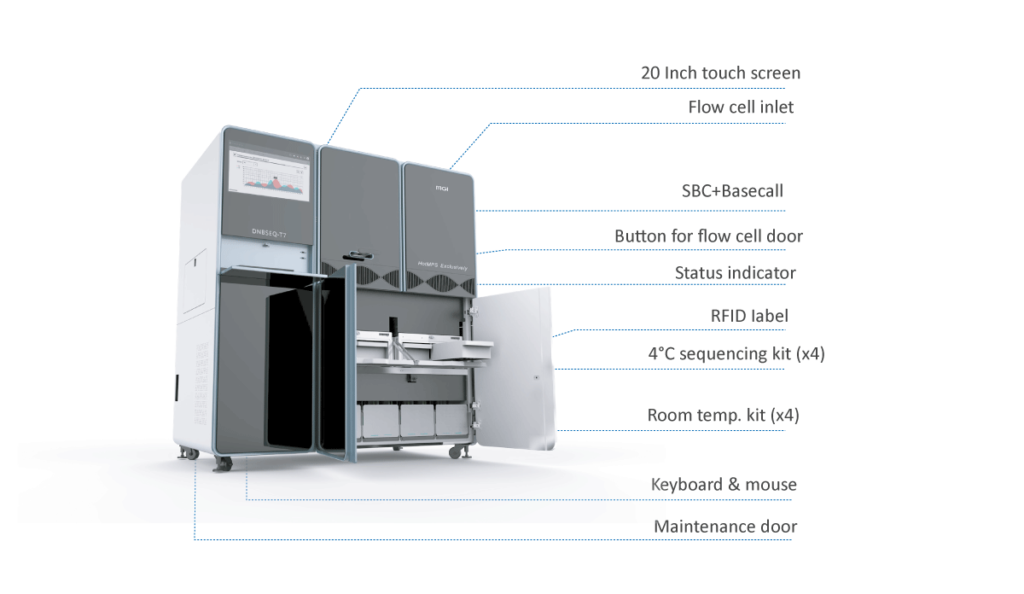
DNBSEQ-T7* can generate 1-4T of high quality data per day, for a wide range of applications including:
- Whole Genome Sequencing
- Deep Exome Sequencing
- Epigenome Sequencing
- Transcriptome Sequencing
- and targeted panel projects.
Powered by DNBSEO™ Technology, DNBSEQ-T7* makes sequencing more efficient and productive with advances in biochemical, fluidics, and optical systems.
PERFORMANCE PARAMETERS DNBSEQ-T7*
1-4 Flow Cells/run, 1 lane/Flow Cell, 5000M max reads/Flow Cell1.
| READ LENGTHS | PE100 |
| DATA OUTPUT | 1-4T |
| Q302 | >85% |
| RUN TIME3 | 20-22 HRS |
- The maximum number of effective reads are based on the sequencing of an internal standard library. Actual output may vary depending on sample type and library preparation method.
- The percentage of base above Q30 is the average of an internal standard library over the entire run. The actual performance is affected by factors such as sample type, Library quality, and insert fragment length.
- Run time includes Flow Cell loading, sequencing, and outputting cal. File. Cal. is a binary file format generated by MGI sequencer basecall software.
More sequencing options for linear library users
The MGIEasy Universal Library Conversion Kit (App-A) Library Compatibility Workflow adapts linear libraries prepared with third-party kits for sequencing on the MGI DNBSEQ™ System. Thereby enabling you to compare or continue your existing workflow on the DNBSEQ™ sequencing platforms.
*StandardMPS sequencing reagents in modified form are available in Germany, UK, Sweden, and Switzerland.
Specifications
| DNBSEQ-T7RS |
for research only |
| Operating Environment Requirements |
Temperature:19~25℃, < 2°C change per hour Relative Humidity:30%RH ~ 80%RH, non-condensing Atmospheric Pressure:80kPa~106kPa Waterproof Rating:IPX0 Altitude:Below 2000 meters |
| Control Computer Configurations |
CPU:Intel CORE I7-7700 4Core*2 3.6GHz Internal Storage:16 GB RAM |
| Power & Dimensions & Net Weight |
Dimensions:Length: 1656 mm, Height: 1815 mm Depth: 903mm Net Weight:765Kg Power Type:200~240V, 50/60Hz, 30A Rated Power:3000VA |
| Bandwidth for Network Connection |
300 MB/s: For local storage network uploads 4000 MB/s: For Fastq computing uploads 500 MB/s: For Data analysis uploads |
